The amazing diversity of proteins encoded by the human genome is expanded by orders of magnitude through post-translational modification (PTM)1,2. Among the most widespread PTMs is long-chain S-acylation. This process is mediated in all eukaryotes by the zinc finger Asp-His-His-Cys motif-containing (ZDHHC) S-acyltransferase family of integral membrane enzymes.
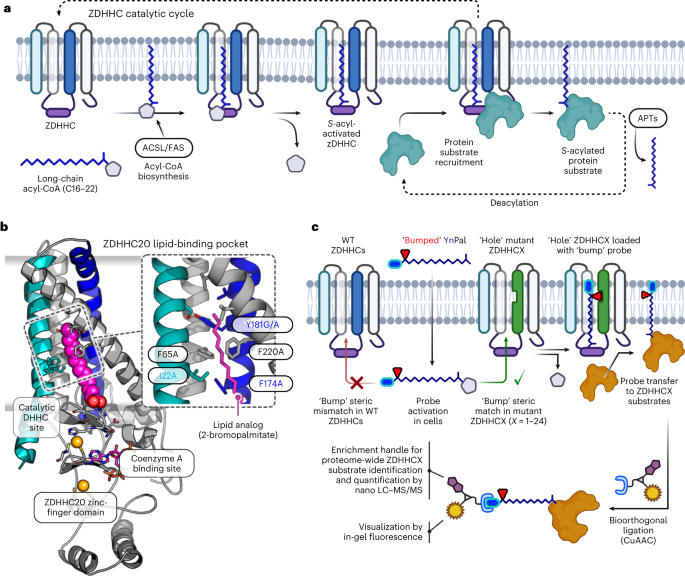
There are 23 known human ZDHHCs that together acylate >3,000 cysteine residues across ca. 12% of the human proteome. The ZDHHC catalytic cycle occurs in two stages: auto-S-acylation of a conserved Cys in the DHHC motif followed by S-acyl transfer to a substrate protein cysteine proximal to the ZDHHC catalytic site.
S-acylation increases local hydrophobicity and membrane affinity and can regulate protein membrane microdomain partitioning, stability, trafficking, nuclear localization, secretion, and protein interactions. De-S-acylation by acyl-protein thioesterase (APT) can generate a dynamic S-acylation cycle implicated in signaling cascades.
During the process, numerous examples of upregulation or downregulation of S–acylation have been found to promote pathological conditions including cancer, inflammatory disease or neurodegeneration.

Despite the importance of ZDHHCs in health and disease, mapping the substrate network of a specific ZDHHC remains a formidable challenge. Global enrichment of S-acylated proteins through metabolic labeling with alkyne-tagged lipid analogs or chemical exchange of S-acyl thioesters for affinity tags can circumvent the difficulty of direct S-acylated peptide detection by liquid chromatography–mass spectrometry (LC–MS), leading to large databases of putative substrate proteins.
However, the lack of selective ZDHHC inhibitors and the confounding influence of ZDHHC overexpression, knockdown or knockout (KO) that can lead to redundancy, compensation or loss of ZDHHC–protein interactions and coregulation currently prevent the direct association of a specific ZDHHC with its cognate S-acylated protein substrates.
Here we establish a chemical–genetic system for direct labeling and identification of the substrates of a specific ZDHHC in intact cells, Read More.
