RNA-binding proteins (RBPs) are involved in various aspects of post-transcriptional gene regulation, including alternative pre-mRNA splicing (AS). Despite the importance of RBPs for cellular function, most of the over 2,000 human proteins that interact with RNA do not have an assigned molecular function. AS plays a crucial role in RNA processing, and aberrant splicing is widespread in diseases such as cancer.
Previous assays have been limited in their ability to identify and characterize RBPs involved in AS. This study aims to address these limitations by developing tethered function luciferase-based splicing reporter assays to systematically assess the modulation of exon inclusion by human RBPs.
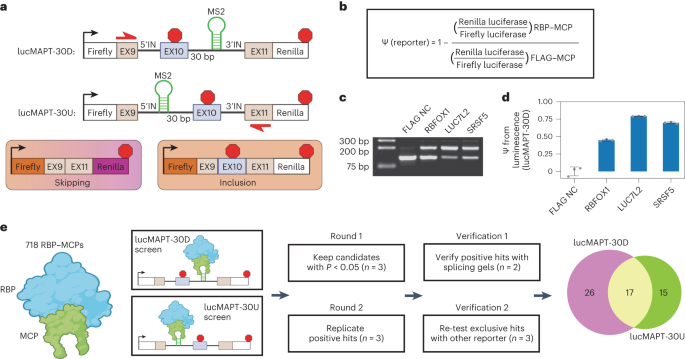
The study constructed two dual-luciferase tethered AS minigene reporter systems based on the splicing event of MAPT (microtubule-associated protein tau) exon 10. This innovative approach allows for the investigation and quantification of the capacity of any protein sequence to directly promote exon inclusion.
By using this system, the study aims to not only reveal factors involved in splicing but also to identify potent and compact exon inclusion activation domains. The assays serve as a biological discovery engine and a prototyping platform for protein engineering applications.
To read more about the study, you can visit the link provided.